Join us for our SDSU TB day on Friday, February 7th, 2020! The event features tuberculosis experts from around the globe to discuss the burden of TB and its relevance to San Diego’s border health.
Author Archive: admin
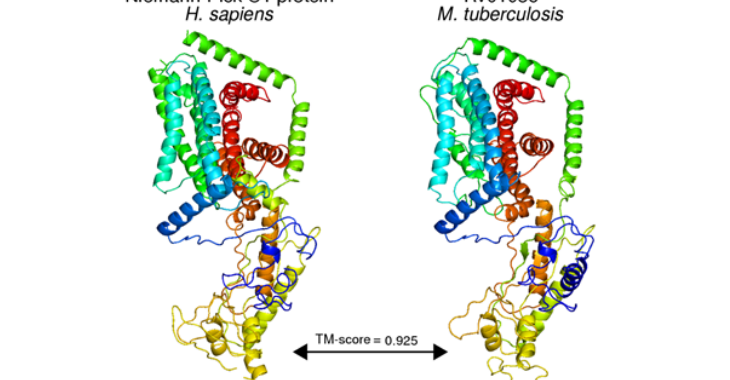
Preprint: SMRT Genome Assembly Corrects Reference Errors, Resolving the Genetic Basis of Virulence in M. tuberculosis
The genetic basis of virulence in Mycobacterium tuberculosis has been investigated through genome comparisons of its virulent (H37Rv) and attenuated (H37Ra) sister strains. Such analysis, however, relies heavily on the accuracy of the sequences. While the H37Rv reference genome has had several corrections to date, that of H37Ra is unmodified since its original publication. Here, we report the assembly and finishing of the H37Ra genome from single-molecule, real-time (SMRT) sequencing. Our assembly reveals that the number of H37Ra-specific variants is less than half of what the Sanger-based H37Ra reference sequence indicates, undermining and, in some cases, invalidating the conclusions of several studies. PE_PPE family genes, which are intractable to commonly-used sequencing platforms because of their repetitive and GC-rich nature, are overrepresented in the set of genes in which all reported H37Ra-specific variants are contradicted. We discuss how our results change the picture of virulence attenuation and the power of SMRT sequencing for producing high-quality reference genomes.
https://doi.org/10.1101/064840
The manuscript is under review by BMC Genomics.
Recent Publications
Valafar lab has new publications! Please check them out
Novel katG mutations causing isoniazid resistance in clinical M. tuberculosis isolates
A Systematic Review of Mutations in Pyrazinamidase Associated with Pyrazinamide Resistance in Mycobacterium tuberculosis Clinical Isolates.
Looking back to ASM 2015
Some of the lab members in Valafar lab attended the ASM 2015 last week in New Orleans. It was great experience for the lab as people were able to catch up with the colleagues around the world and catch up with latest trend in the field of microbiology. The lab was able to share information acquired during the convention and held a meeting to discuss where to head from here. We would like to thank ASM for the great event and we look forward to going again next year.
New Website!
This post will be replaced whenever new posts are added!!